OdontoTest: Genetics and microbiology in oral diseases
Test to analyse genetic variants and types of bacteria that influence the occurrence and development of periodontitis, peri-implantitis and caries.
Price
On request, depending on the test
Time to result
On request, depending on the test
What is the OdontoTest for?
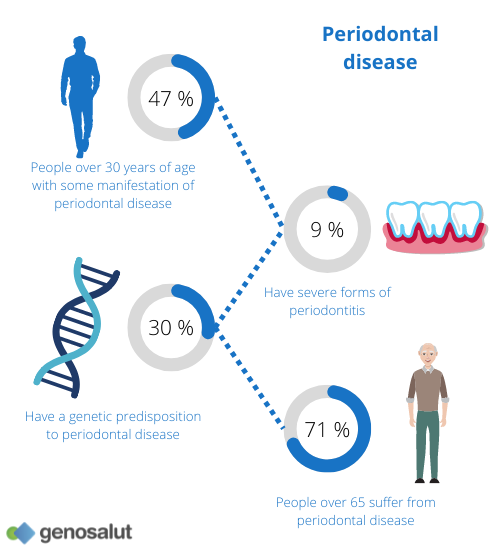
The OdontoTest is used for the detection of relevant microorganisms involved in the development of periodontitis, peri-implantitis and caries as well as for the genotyping of the most significant gene variants related to individual susceptibility to the development of these disorders.
Caries and periodontitis are the most common infectious diseases of the oral cavity. Caries is related to the destruction of dental hard tissues, while periodontitis affects the soft tissues and alveolar bone. Gingivitis is a reversible condition that affects the gums, but may favour the development of periodontitis.
These diseases are classically associated with dysbiosis of the commensal microbial flora of the oral cavity. However, in the current concept of oral cavity infectogenomics, host genetic factors are also considered relevant and may exert a significant influence on the formation and composition of microbial biofilms, as well as on the immune response to this microbial challenge.
What does the OdontoTest analyse?
Using a saliva sample obtained by rinsing the mouth with sterile saline, we analysed the presence of periodontopathogenic and cariopathogenic bacteria and the genetic variants associated with these conditions by qPCR-HRM.
Pathogenic bacteria
qPCR using the modified Pfaffl relative quantification model for counting micro-organisms in ug/μl for the detection of the following bacteria:
- S. mutans
- Lactobacillus spp
- Actinomyces spp
- A. actinomycetemcomitans
- P. gingivalis
- T. forsythia
- T. denticola
- P. micra
- P. intermedia
- F. nucleatum
Genetic variants
The qPCR-HRM technique is applied to detect variants in the following genes:
- MMP16
- AMELX
- DEFB1
- GLT6D1
- IL-1B
Our value proposal
Experience
At Genosalut, we have more than 10 years of experience in counselling people with conditions where a genetic cause has been identified or is thought to be possible.
Proximity
We are a close laboratory, we respond personally and we take the time to explain the report in detail to doctors and patients.
Professional interpretation of results
Because of our knowledge and experience, we are able to accurately interpret genetic results and offer professional advice.
Reference in the field
We are the contact for patients, dentists and clinics in all areas of genetic and microbiological diagnostics and prevention.
Who is the OdontoTest aimed at?
Genetic and microbiological studies for dentists offer solutions and support in the prevention, diagnosis and treatment of oral diseases such as periodontitis, peri-implantitis or caries. They are especially recommended in the following cases:
Periodontitis
In cases of: refractory and therapy-resistant adult periodontitis, rapidly progressive acute periodontitis and periodontal disease with pocket depth > 4 mm (despite optimal oral hygiene).
Implantology
In cases of peri-implantitis and before each implant.
Caries
In cases of people with a high predisposition to develop caries.
Information from the OdontoTest
Prevention
It allows the dentist to determine the best measures to prevent the onset and development of these pathologies and to give individualised advice to his patients.
Optimisation of the therapeutic strategy
Knowledge of bacteria can help to improve antibiotic treatment. This knowledge combined with knowledge of genetic variants can, for example, help determine the likelihood of implant success.
Determination of the follow-up examinations
To optimise the operation of the practice, while minimising the inconvenience to patients due to unnecessary travel.
Personalised medicine
To offer a quality service based on the individual needs of each patient.
How can I request a OdontoTest
Request appointment
Contact us through the form, by e-mail or by telephone to make an appointment with us.
We analyse the probe
In our genetic diagnostics laboratory we analyse the sample with the latest technology.
We write a report
We provide a detailed description of the results and, if necessary, genetic counselling.
FAQs
What is the oral microbial flora and what is its function?
The microbial flora or oral microbiota is made up of a set of microorganisms that form a true microecosystem in the oral cavity. Its functions include protecting the host against infections caused by colonisation of the oral cavity by pathogenic microorganisms from the outside and from other cavities such as the ear and nose, as well as stimulating the immune system and keeping the mucous membranes in good condition.
What does the composition of the oral microbial flora depend on?
The composition, abundance and diversity of microbial species in the oral flora depends on many factors.
- During the first years of life, the main route of transmission of micro-organisms in the oral cavity is from mother to child through direct contact and breastfeeding (vertical transmission).
- Subsequently, the route of transmission through direct contact with other family members, including the father, siblings and other potential caregivers (horizontal transmission), as well as other environmental factors, such as diet, oral hygiene, antibiotic intake and smoking, becomes important.
The variability is so great that no two people have completely identical oral microbial flora.
How many microorganisms make up the oral microbiota?
To date, the Human Oral Microbiome Project has identified more than 700 microbial species in the oral cavity, with the generaNeisseria, Spreptococcus, Actinomyces, Lactobacillus, Veillonella and Granulicatella being the most representative in quantitative terms.
What are the periodontopathogenic and cariopathogenic bacteria?
Less than 1% of the species identified in the oral microbiota are involved in the development of caries and periodontitis. An imbalance in the normal microbial populations of the oral cavity may favour the overgrowth of this small group of aciduric and acidogenic microorganisms with pathogenic potential.
- The following microbial agents are involved in the pathogenesis of caries: Streptococcus mutans, Lactobacillus spp. and Actinomyces spp. Of these microorganisms Streptococcus mutans has the highest cariogenic potential.
- The pathogenesis of periodontitis is due to other micro-organisms classified into two complexes, according to the degree of pathogenicity.
- The red complex includes three of the most pathogenic periodontal bacteria: Tannerella forsythia, Treponema denticola and Porphyromonas gingivalis.
- The second group of periodontal pathogenic species is known as the orange complex and includes Prevotella intermedia, Fusobacterium nucleatum and Parvimonas micra.
- Another microorganism with great periodontopathogenic potential is Aggregatibacter actinomycetemcomitans, which acts as an immunosuppressive agent of the host periodontal defence system and alveolar bone homeostasis.
Genetic variants associated with oral diseases: inclusion criteria
Based on the phenotype-SNPs association provided by the SNPedia database, the recommendations of the American Dental Association and the literature sources consulted, a total of five variants assessing individual susceptibility to the development of caries and periodontitis have been selected, prioritising those that have been previously identified by GWAS (genome-wide association studies) and subsequently validated by independent trials or meta-analysis studies. These variants are as follows:
- rs1047031 (GLT6D1)
- rs1047031 (DEFB1)
- rs2046315 (MMP16)
- rs17878486 (AMELX)
- rs1143634 (IL1B)
These are mainly genes that play a role in the composition of the tooth and the inflammatory response of the host. The presence of any of these variants per se does not imply that the carrier will develop caries or periodontitis, as both conditions are the result of a multifactorial process with several factors involved.
Gene GLT6D1
The rs1047031 variant is located in GLT6D1, a gene with a high expression rate in gums. The exact function of this gene is currently unknown. However, the C allele of this gene variant has been associated with the development of severe periodontitis in different populations.
Gene DEFB1
The rs1047031 variant is located in the regulatory region of the DEFB1 gene, which encodes a defensin family protein with broad-spectrum antimicrobial activity. The DEFB1 gene appears to play an important role in the maintenance of oral health. The T allele of rs1047031 has been associated with chronic and severe periodontitis, in addition to other oral cavity pathologies such as caries. The risk is higher when the homozygous recessive genotype for this allele (TT genotype) is present.
Gene MMP16
The rs2046315 variant is located in the regulatory region of the MMP16 gene. This gene encodes a protein of the metalloprotein family that participates in the degradation of extracellular proteins in multiple physiological processes and plays an important role in cariogenic processes. In fact, the T allele of this variant has been associated with an increased risk of caries development.
Gene AMELX
The rs17878486 variant is located in AMELX, a gene encoding a prostaglandin that plays a very important role in the development of tooth enamel. The T allele of this variant has been associated with an increased risk of caries, with the TT and TC genotypes being the most at risk. However, this allele has a high allele frequency in the European population (around 85%).
Gene ILB1
The rs1143634 variant is located in the IL1B gene, which belongs to the interleukin family, and is involved in several processes necessary to initiate and maintain the inflammatory response. Altered IL1Bla protein values have been found in patients carrying the T allele of this gene variant with periodontitis. This allele has been associated with chronic periodontitis, with carriers of this allele (TT and TC genotypes) having an increased risk of periodontitis.
Request an appointment with us
Opening hours
Monday to Friday from 9.00 am to 1.00 pm
+34 616 59 01 65
info@genosalut.com
Camí dels Reis, 308 (Clínica Palma Planas)
Contact form
Reasons for trusting Genosalut
First genetic diagnosis laboratory in the Balearic Islands
Professionals with experience in medical genetics
Detailed report of the results
Personalised attention for each patient
Wide range of genetic tests
Cutting-edge technology
